
Whether you’re a seasoned computational scientist or an armchair immunologist, the growing focus on the T-cell triad — MHC, peptide & TCR — in both infectious and chronic diseases is necessitating easy access to complex datasets and analysis tools that can make sense of the human adaptive immune response.
In a recent webinar, DNAnexus research platforms expert Ben Busby joined Ankita Das, Head of Product at immune profiling company MIODx, to discuss ways in which the companies are working together to make it easier to interrogate and interpret TCR (T-cell receptor) data.
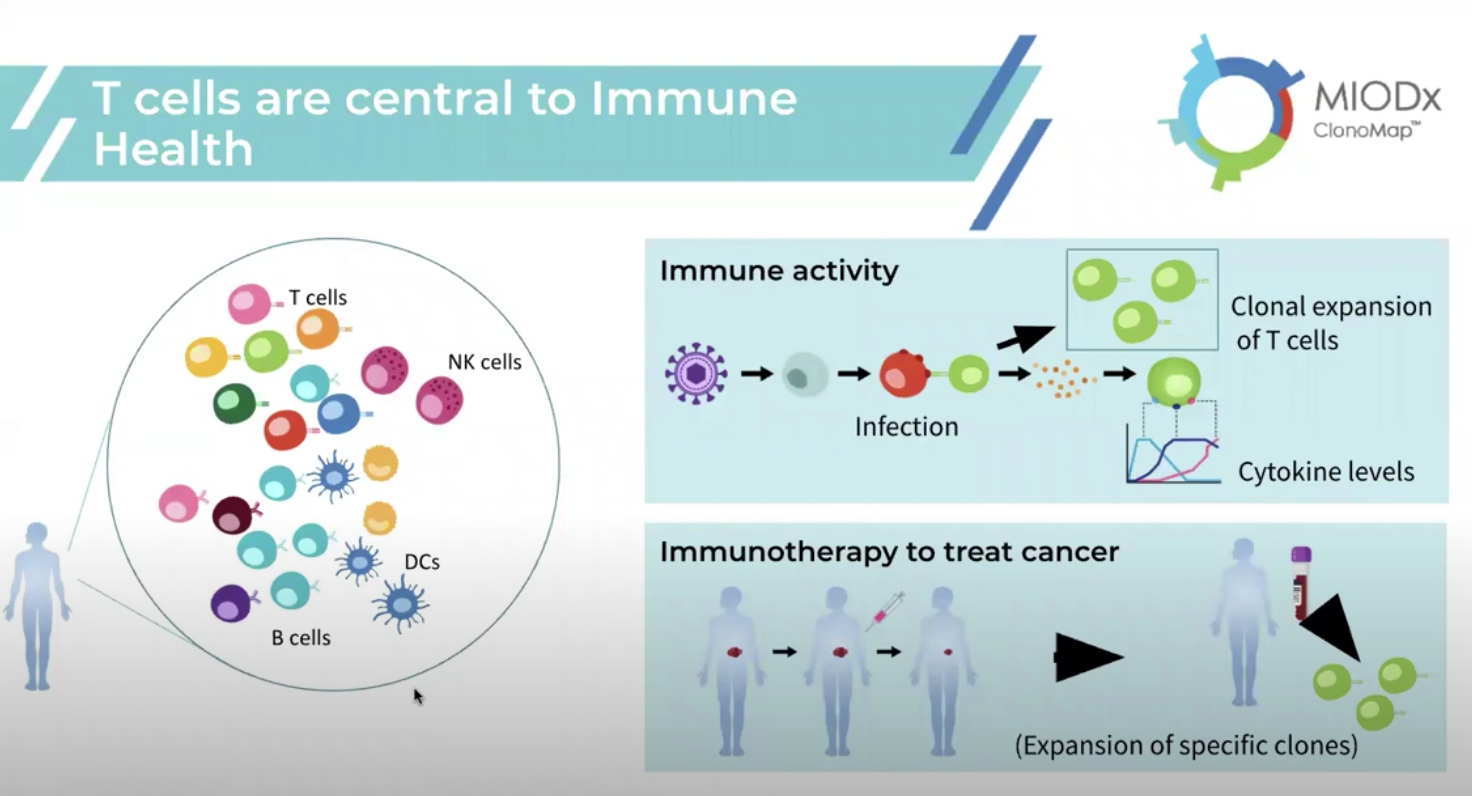
As Das explained, T-cell composition and activity are at the center of the immune response and key to tracking immune health. The composition of a person’s T-cell repertoire can vary depending upon factors like age, environment, genetics, infection, and lifestyle. When T-cells sense an infection, they undergo a phenomenon called clonal amplification, wherein a subset of the T-cell repertoire will amplify to orchestrate an immune response, whether that be killing off infected cells or recruiting B cells to generate antibodies. Interpreting clone activity can reveal important clues about immune health and insight that could potentially lead to biomarker and therapeutic discovery.
However, scientists wishing to undertake such research face several challenges. “TCR data is currently very siloed, and the architecture available for hosting and analyzing the immense data sets involved are often not scalable,” Das said.
The MIODx team set out to overcome these challenges by creating ClonoMap™, a SaaSportal hosted on DNAnexus Titan™, in which TCR repertoire libraries can be stored, managed and analyzed in a secure environment.
ClonoMapTM subscribers upload raw sequence data from TCR repertoire libraries into the portal and run the ClonoMap™ Immune Profiler analysis to discover repertoire features. A second tool, ClonoMapTM Immune Insight, searches public datasets to help scientists see the new immune profiles in context and draw translational insights, such as biomarker identification. The MIODx team is now taking it a step further, applying machine learning to create personalized, immune health scores.
The ClonoMapTM suite of tools has already received nods for its innovation during the precisionFDA COVID-19 Precision Immunology App-a-thon. In the webinar, Das shared examples of how it was used to generate data and insights in COVID-19 and rheumatoid arthritis. In the case of COVID-19, for example, the Immune Profiler highlighted specific T Cell Receptor Beta Variable (TRBV) genes and CDR3 clonotypes at different frequencies in healthy individuals compared to COVID-19 recovered patients, providing TCRs for further investigation with respect to disease severity.
Holy grail of healthcare
Unravelling the immune response is the ‘holy grail of healthcare,’ Busby said. And in what may prove to be the decade of infectious disease research, Busby said he was proud to be able to provide tools to help scientists do so.
“We want to make this size data accessible to everyone, and the DNAnexus platform really enables scientists and bioinformaticians to be more powerful,” Busby said.
In addition to ClonoMap™, the iReceptor data discovery platform, curated by the AIRR (Adaptive Immune Receptor Repertoire) community, facilitates the curation, analysis and sharing of antibody/B-cell and T-cell receptor repertoires (AIRR-seq data) from multiple labs and institutions.
JupyterLab is another powerful tool that DNAnexus leverages for multi-omic cohort analysis and data exploration, and Busby recommended it for experts and armchair enthusiasts alike. He also noted that many DNAnexus-created Jupyter Notebooks are available, even to non-DNAnexus users.
Other open source tools include Bioconductor, Bioconda and Docker. Each can be easily integrated into DNAnexus platforms, and the visualization capabilities of the DNAnexus system make the data obtained by them even more approachable, Busby said.
You can watch the full webinar below.

.png)
.png)
.png)