Four years ago, precisionFDA, in collaboration with The Genome in a Bottle (GIAB) Consortium led by the National Institute of Standards and Technology (NIST), successfully completed the Truth Challenge for human germline variant calling using a reference material HG002, also known as NA24385, the son of an Ashkenazi trio. This resulted in significant advances in variant calling accuracy, and enrichment of well-characterized benchmark datasets on precisionFDA with contributions from Illumina, Garvan, Macrogen, and NIST. The original challenge was timed such that a GIAB benchmark for reference material HG001 had been publicly released, but the benchmark for HG002, while complete, had not been made public. The HG002 benchmark was thus used as the blind truth set by which challenge submissions were graded based upon their results concordance with the HG002 benchmark produced by GIAB.
Recently, new long and linked read technologies and bioinformatics pipelines have enabled the characterization of increasingly challenging regions of the genome, such as an expanded benchmark developed by the GIAB for the parents of HG002 (HG003 and HG004). Another such propitious timing of benchmark releases has presented the opportunity for the Truth V2 Challenge, focusing on variant calling in difficult-to-call genomic regions in GRCh38. While HG002 Benchmark V4.1 has been publicly released, the recently completed benchmarks for the parents of HG002, namely HG003 and HG004, have not yet been made public, and will be used as the blind truth set. This challenge aims to provide a common frame of reference for measuring performance aspects of participants’ pipelines on difficult-to-map regions, segmental duplications, and the Major Histocompatibility Complex.
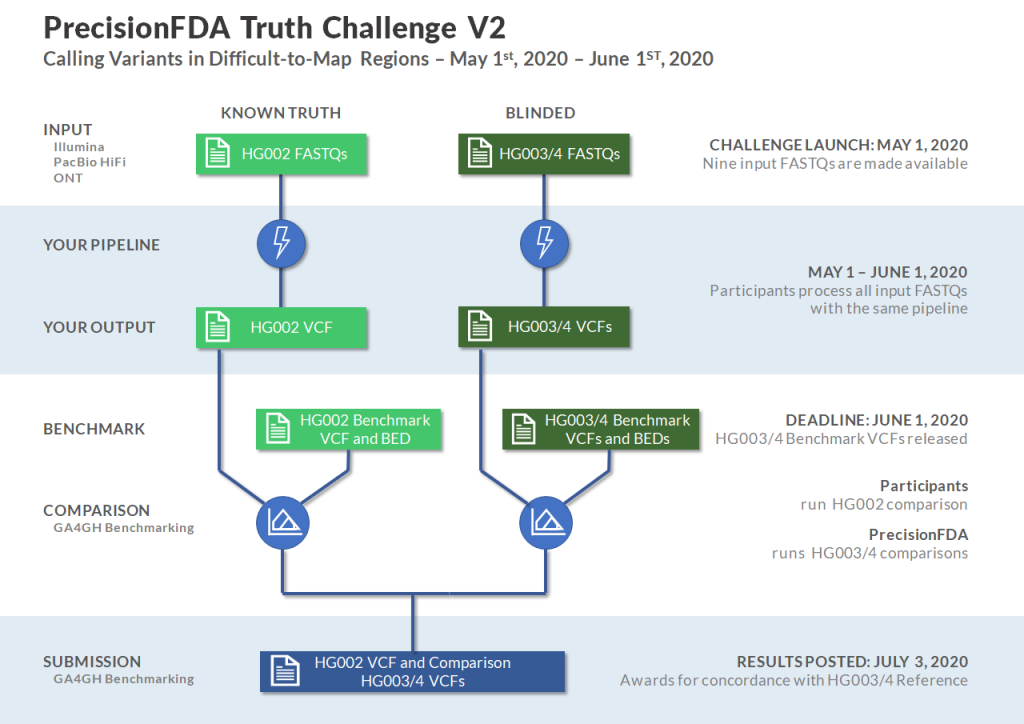
Participants in the Truth V2 Challenge will be provided nine FASTQ files, three datasets for each of HG002, HG003, and HG004, produced by Illumina, PacBio HiFi, and Oxford Nanopore sequencing technologies. Participants will also be provided with the benchmark VCF and BED files for HG002 with which they can compare the results of their pipeline using the GA4GH Benchmark Comparator on precisionFDA. Submissions will include VCF files for HG002, HG003, and HG004, and the HG002 benchmark comparison results. Submissions will be graded on sensitivity and specificity independently and awards will be presented on precisionFDA for best concordance with the HG003 and HG004 Benchmarks which will be published simultaneously with the closing of the challenge.
The Truth V2 Challenge begins May 1, 2020 with a submission deadline of June 1, 2020. To pre-register for the challenge, visit our sign up!

.png)
.png)
.png)