The application of next-generation sequencing in the study of microbial communities has fueled the rapid growth of interest in microbiome research. However, difficulties with the accuracy of computational analyses of these complex datasets have limited the translation of microbiome science into novel biotherapeutic products. In order to unleash the potential that metagenomics holds for human health, computational methods to identify unique microbial strains must be improved.
The Mosaic Community Challenge: Strains #1, sponsored by the Janssen Research & Development, LLC, through the Janssen Human Microbiome Institute, aims to benchmark and improve the performance of computational tools in analyzing these data, in order to provide better quality profiling of microbiome samples at high resolution. The challenge gave participants the opportunity to validate their bioinformatics tools in realtime on a neutral, unbiased platform, and see how they performed against other industry tools.
Participants of the challenge worked with datasets that were composed of four different sample types: a metagenomics dataset generated from real mouse fecal samples (of known bacterial composition), and three simulated datasets of varying complexity. Besides the challenge dataset, a distinct training dataset, which included the truth files, was provided to enable participants to train and improve their methods. Participants were then able to conduct analysis by either creating their own app on the Mosaic Platform, or by downloading the dataset and running their method in their own system. Over the four-month course of the challenge, participants could take advantage of a “Testing Ground” to get immediate feedback on their work with training datasets before submitting their final challenge entries.
Challenge Winners & Their Methods
We would like to congratulate the winners as well as thank all who participated for helping to take microbiome science to the next level.
Profiling
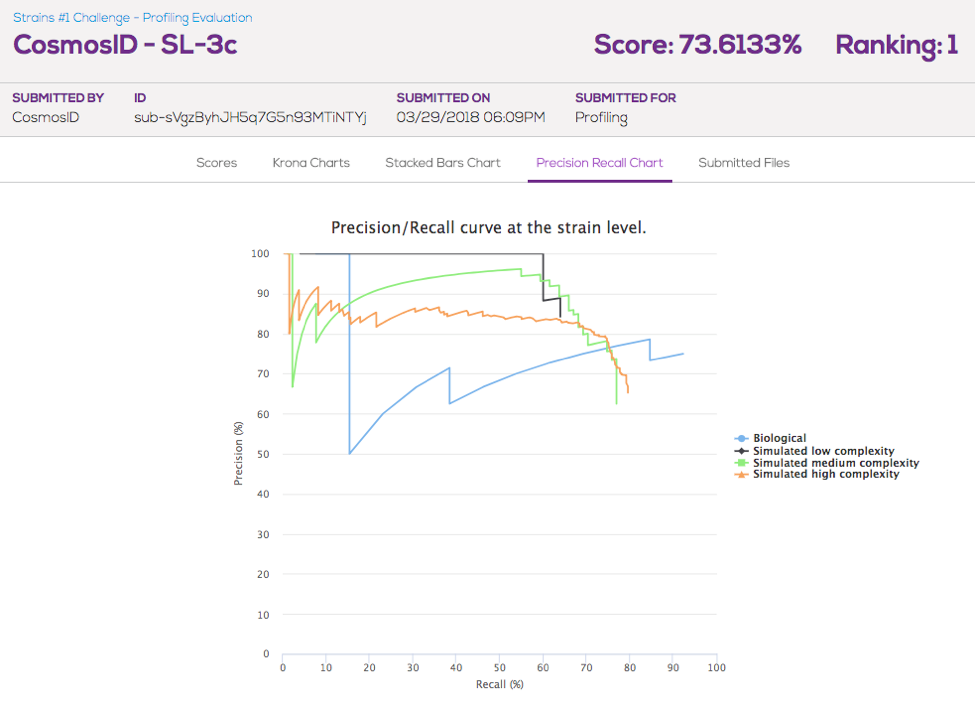
CosmosID, a bioinformatics and NGS service laboratory, scored highest in the Profiling part of the challenge. The CosmosID analysis pipeline achieved the highest cumulative F1-score, which is a measure of precision and recall. According to Nur Hasan, Chief Science Officer at CosmosID, the strength of their approach lies with the manually curated database, whose structure follows the phylogenetic hierarchy of all represented microorganisms which enables reliable microbial identification at all taxonomic levels, down to strain-level.
CosmosID’s submission scored the highest in the analysis of the Biological Sample (80%), which was 64% higher than the score of the second submission (48.9%). Interestingly, however, submissions based on the popular Metaphlan tool, performed better across the simulated datasets. The observation that the performance of tools vary based on the source of the sequencing data highlights the importance of benchmarking the tools on both biological and simulated datasets.
Figure 1. Precision/Recall Curve for the winning submission for each of the challenge datasets (to view this chart visit the submissions page on Mosaic).
To interactively compare the Profiling submissions and view Precision Recall Curves, visit the Strains #1 Profiling comparison page.
Assembly
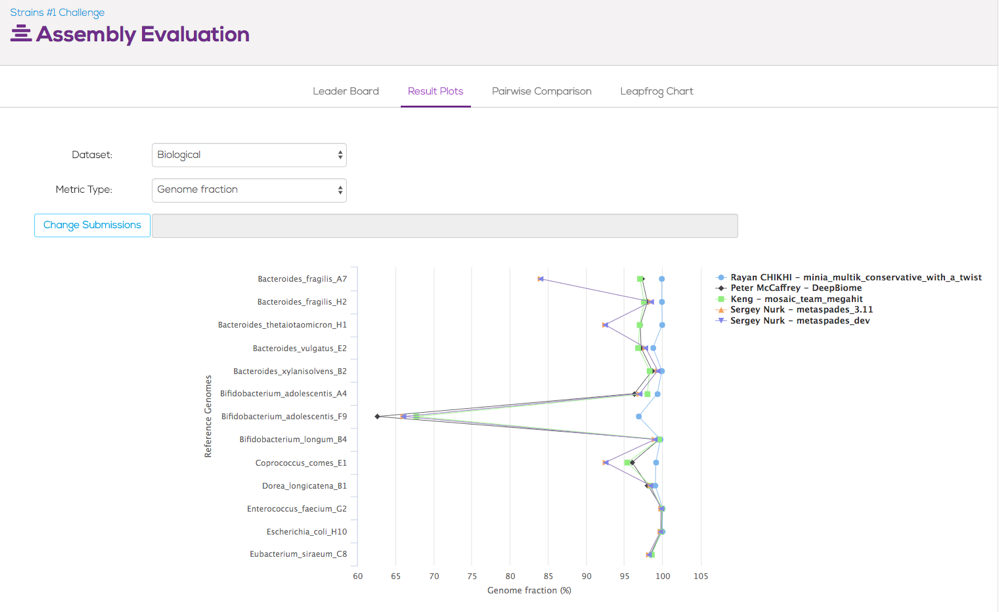
Rayan Chikhi, PhD, Computer Scientist at the French National Center for Scientific Research (CNRS) and CRIStAL research center, and an advisor at Clarity Genomics, scored highest in the Assembly part of the challenge by using the Minia assembler to assemble the metagenomic data provided for the challenge. The assembly portion was judged on the total number of aligned bases divided by the reference genome size (Genome Fraction). The winning submission scored well across all other metrics reported in the leaderboard, namely Misassemblies and Mismatches.
Figure 2. Genome fraction scores across 13 biological sample reference strains
Honorable mentions go to two other participants. Peter McCaffrey came a close second with his DeepBiome submission, while his submitted assemblies were longer than the winning submissions. Additionally, the submissions from Sergey Nurk (Metaspades assembler) had consistently the largest contigs.
To make your own comparisons between the submissions and dive in deeper in the rich comparison data available, visit the Strains #1 Assembly comparison page.
Learn about the winners’ methods during our webinar confirmed for Tuesday, June 26th at 10am PT (1pm ET).
Want More Ways to Participate in the Mosaic Microbiome Community?
Learn more and get involved at mosaicbiome.com/challenges.
Visit Us at Microbiome Drug Development Summit!
DNAnexus will present Translation of Microbiome Research into Clinical Applications, this Friday, June 22nd at 12pm at the Microbiome Drug Development Summit in Boston. Join our talk, and stop by our exhibition table to learn more about DNAnexus microbiome capabilities, and the Mosaic Community Platform & Challenges. Email us to schedule a meeting in advance.
Translation of Microbiome Research into Clinical Applications
- Crowdsourcing the advancement of microbiome research with the Mosaic Community platform and challenges
- Considerations for incorporating microbiome data into clinical trials
- Complying with GLP, 21 CFR Part 11, and more
Speakers:




.png)
.png)
.png)