Our mission, and we chose to accept it, was to join more than 100 researchers and engineers to look for answers and create insights into a real patient’s mystery medical condition.
Our mission, and we chose to accept it, was to join more than 100 researchers and engineers to look for answers and create insights into a real patient’s mystery medical condition.
In this case, that patient was John, aka “Undiagnosed-1”, a 33-year-old Caucasian male suffering from undiagnosed disease(s) with gastrointestinal symptoms that started from birth. In his 20’s, John’s GI issues became more severe as he began to have daily lower abdominal pain characterized by burning and nausea. He developed chronic vomiting, sometimes as often as 5 times per day. Now 5’10” tall and 109lbs, he is easily fatigued due to his limited muscle mass and low weight.
Armed with more than 350 pages of PDFs containing scanned images of John’s medical records plus a range of genetic data — from Invitae’s testing panel to whole genome shotgun sequencing from multiple technologies (Illumina, Oxford Nanopore and PacBio) — could we generate ideas for diagnosis, new treatment options or symptom management? Perhaps we could interpret variants of unknown significance or identify off-label therapeutics through mutational homogeneity.
This was the challenge set to us as part of a three-day event in June organized by SV.AI, a non-profit community designed to bring together bright minds from AI, machine learning and biology backgrounds to solve real-world problems. This was its third event. Last year, we helped to apply DeepVariant from Google on a new kind of sequencing data to help out on a rare kidney cancer case.
Off to a flying start
At DNAnexus, our main mission is to help our customers to process large amounts of genomic data with cloud computing. It is straightforward for us to do the initial processing of the genomic data. In this case, our customers were the event’s community of genomic “hackers.” We decided to pre-process John’s genomic data, so that our fellow participants would not have to spend extra effort to go through variant calling.
Chai generated SNPs and structural variants before the event, and made the information available to everyone who might need it.
However, genomic data was only half of the picture. Clearly, the clinical information gleaned from John’s medical records would shed some clues. But how to get through hundreds of pages of scanned images of his medical records (under MIT licence for invited participating scientists). There must be a smarter way to process the records that would allow us — and others — to write scripts and programs to process them.
Luckily, there is: Amazon Textract and Amazon Comprehend Medical.
Developed by Amazon Web Services (AWS) using modern machine learning technology, Amazon Textract is a service that automatically extracts text and data from scanned document — think OCR on steroids. While there are many OCR software applications on the market these days, Textract provides more; it detects a document’s layout and the key elements on the page, understands the data relationships in any embedded form or table, and extracts everything with its context intact. This makes it possible for a developer to write code to further process the output data in text or JSON format to extract important information more efficiently. It is important to note that at the time of this blog post, Amazon Textract is not a HIPAA eligible service. We were able to use it in this case because the patient data being analyzed was de-identified. Amazon Textract should not be used to process documents with protected health information until it has achieved HIPAA eligibility. Please check here to determine if Amazon Textract is HIPAA eligible.
Another technology developed by AWS, Amazon Comprehend Medical, was used to process the output from Textract for John’s medical records. Amazon Comprehend Medical is a natural language processing service that makes it easy to use machine learning to extract relevant medical information from unstructured text. Using Amazon Comprehend Medical, you can quickly and accurately gather information, such as medical condition, medication, dosage, strength, and frequency from a variety of sources like doctors’ notes, clinical trial reports, and patient health records. Using it, we were able to extract medications, historical symptoms, and medical conditions from John’s doctors’ notes and testing/diagnostics reports.
With more structured patient information than merely a collection of medical records as images, combined with the variant calls generated from the genomics data, the participants of the hackathon were able to jump right into solving the medical and genomic puzzles for John to help him and relieve his symptoms.
The results
We were happy that a couple of teams from the hackathon were able to use both the variant call set and the processed medical data that we provided.
- The “Too Big to Fail John” team was able to develop an approach to combine the medical conditions and the genomics information to generate a list of potential important variants and apply pathway analysis.
 A nice word cloud generated from John’s medical record and genomics information by the “Too Big to Fail John” team. (Origin: https://github.com/SVAI/Undiagnosed-1/tree/master/TooBigToFail)
A nice word cloud generated from John’s medical record and genomics information by the “Too Big to Fail John” team. (Origin: https://github.com/SVAI/Undiagnosed-1/tree/master/TooBigToFail)
- The “Thrive” team used a similar approach to find potential variant candidates by identifying variants that are commonly seen with John’s medical conditions and predicted deleterious variant calls.
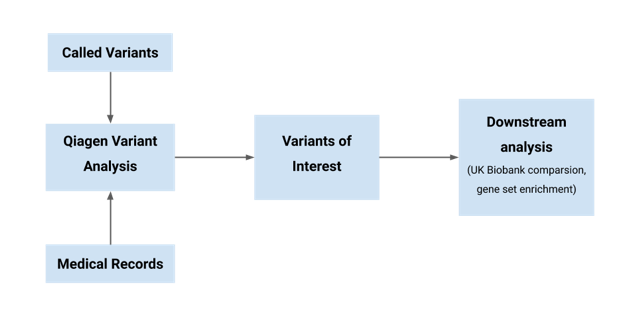
The schema of the Thrive team approach to analyzing both the medical records and variant call set.
- The team Crucigrama extended the scope by including other public ‘omics data, such as metabolic profiles and NLP process, for public genomics data.
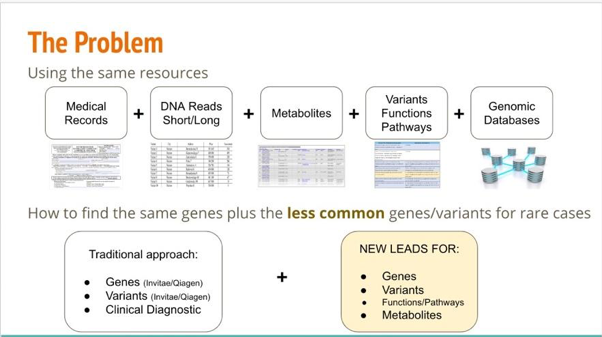 Team Crucigrama’s problem statement to extend the traditional approach to finding new leads for Undiagnosed-1.
Team Crucigrama’s problem statement to extend the traditional approach to finding new leads for Undiagnosed-1.
- The “Beyond Undiagnosed” team also utilized the medical record in their workflow so they could gather key symptoms and diagnoses fast, to provide future care recommendations according to their findings.
 The “Beyond Undiagnosed” team used the extracted symptoms from the medical notes in their workflow for providing recommendations. All information was de-identified and did not contain PHI.
The “Beyond Undiagnosed” team used the extracted symptoms from the medical notes in their workflow for providing recommendations. All information was de-identified and did not contain PHI.
We found it very inspiring that John, who attended the event, was willing to share his medical records with all of the participants in order to help us help him — and we hope the work we did will ultimately do so.
For more information about John’s case, visit the SV.AI site, which will provide all data so that any researchers can continue working on it. We also like to thank SV.AI organizing this event and Mark Weiler and Lee Black from Amazon helping us on processing the data through Amazon Textract and Amazon Comprehend Medical.